table of contents
Digital PCR as a robust tool for low-level pathogen detection

Reliable and accurate quantification of pathogens is essential for determining (low) microbial levels in complex biological samples. However, the accuracy of culture‑based and molecular methods can differ, particularly at low target concentrations or in the presence of inhibitory substances from complex matrices such as feces or cloacal swabs. A direct comparison of these methods is therefore needed to identify which technique provides the most reliable and accurate results.
How We Investigated the Problem
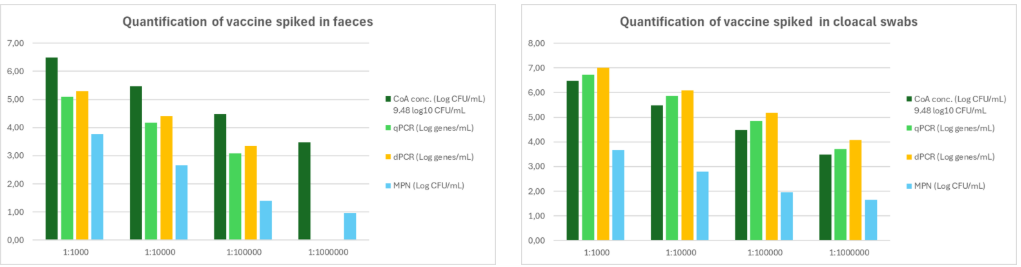
A 1:10 dilution series was prepared from a Salmonella vial with a known concentration (9.48 log10 CFU/mL). This 1:10 dilution was used to compare four quantification methods: (1) classical bacterial enumeration, (2) Most Probable Number (MPN), (3) quantitative PCR (qPCR), and (4) digital PCR (dPCR). The same dilution was also used to spike fecal material and cloacal swabs, enabling evaluation of potential matrix effects for each method.

Results
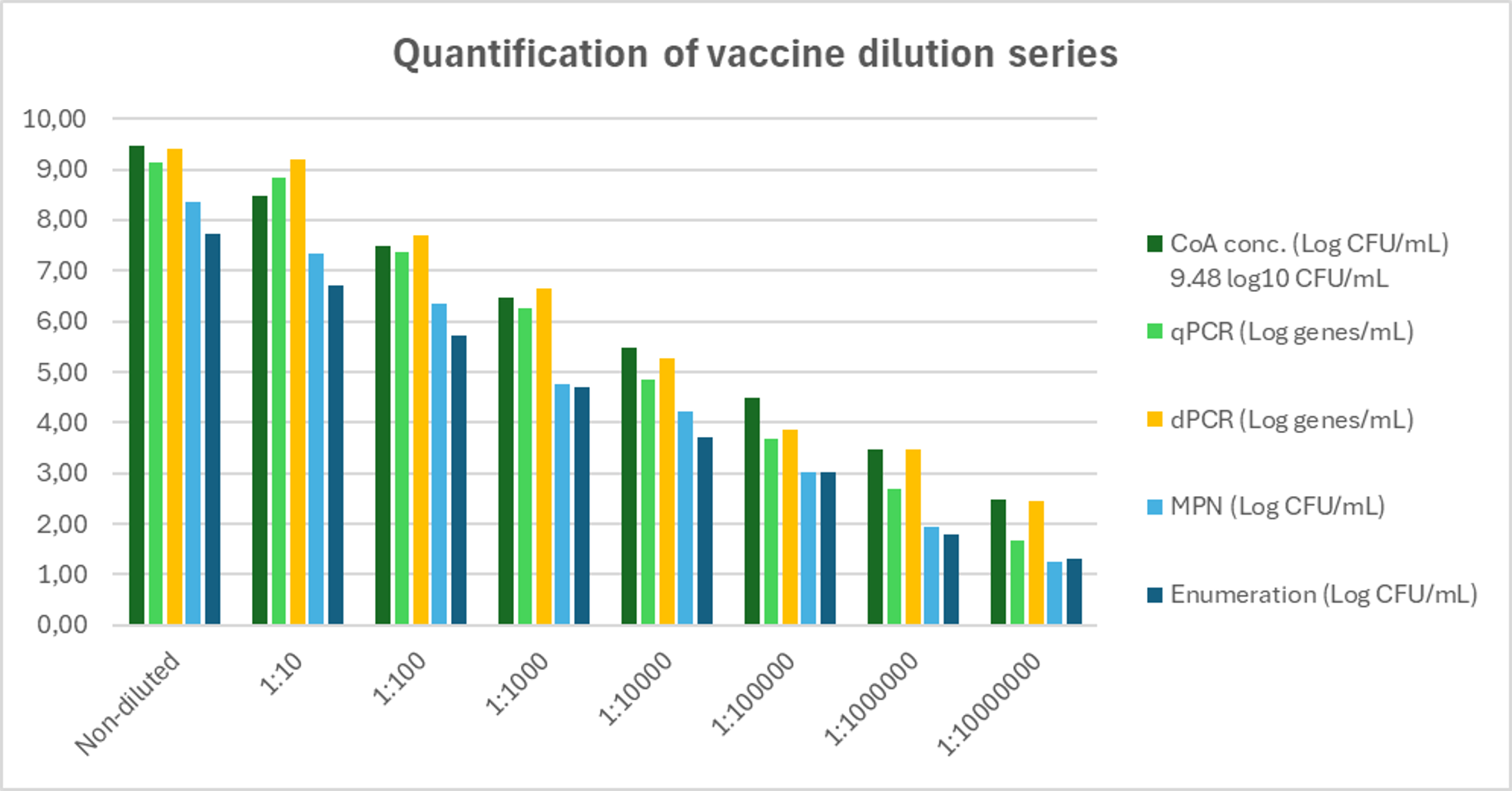
Quantification of the vaccine dilution series
When comparing the four different methods, both molecular methods matched the expected concentrations in the higher dilution range (log9–log6). At lower concentrations (log5–log2), qPCR showed a slight loss in quantification, whereas dPCR continued to produce the expected values. The two bacterial methods showed a consistent pattern across all dilutions, measuring approximately one log lower than expected.